The Arizona Genetics Core now offers High Throughput sequencing using the Element AVITI System. Supported applications include DNASeq, RNAseq, targeted gene sequencing, Metagenomics (16S/custom), small genome sequencing, amplicon sequencing and other NGS pipelines.

General Information
Sample Input
DNA or RNA from a variety of sample types can be run on the AVITI System including cells, tissue, and blood. We also offer DNA or RNA extraction for library prep.
Applications include:
- Small Genome Sequencing (Shotgun/de novo)
- Metagenome Analysis (16S/custom)
- CHiPseq
- miRNA profiling
- Cancer Genomics (RUO)
- RNASeq (transcriptome sequencing/expression analysis)
- Single Cell sequencing
- Other supported and custom protocols (please inquire)
Flexible Outputs
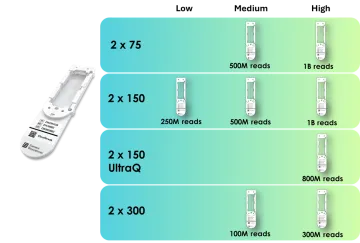
The Element AVITI offers several flow cell read lengths and sequencing outputs to ensure the best fit for your project needs. Our core can help you select the best flow cell for your project.
Data is now returned via Cyverse. This platform offers storage and data analysis tools.
https://learning.cyverse.org/account/
https://learning.cyverse.org/ds/move_data/
Register for an account once your sequencing order is complete.
Data delivered is Fastq format
Additional data analysis is available on request.
Details
Sample Preparation Details
DNA Sample Requirements (if not submitting tissue, cells, etc. for extraction):
- DNA should contain no particulate matter
- DNA must be double stranded
- DNA should not be degraded
- DNA sample should be suspended in LowTE or water
RNA Sample Requirements:
- RNA should be resuspended in RNAase free water and kept at -80°C until delivery to Arizona Genetics Core
- RNA should not be degraded
Workflow
Complete the Request a Quote form and we will contact you to discuss your project workflow.
Quality Control of Samples: Every sample received for sequencing will go through a set of quality control checks before it can be processed. Customers may be asked for more DNA/RNA if their sample fails either of the following checkpoints:
- Pico/RiboGreen Assay to verify the quantity of starting material
- Bioanalyzer DNA/RNAchip to check the material quality
Sequencing Chemistry: The AVITI Stystem uses Avidity base chemistry (ABC). The fundamentals of ABC technology leverage the unique properties of avidites to execute an efficient sequencing reaction and yield highly accurate data. A strong signal-to-noise ratio that persists through high polony densities drives this accuracy. When a run starts, the library hybridizes to surface primers coating the flow cell. Amplification polymerase then binds to the library and primer duplexes, catalyzing rolling circle amplification (RCA) and generating long DNA strands that include copies of the original library ). Each strand forms a polony that contains hundreds of copies of the original library. The polonies hybridize to read-specific sequencing primers. For each cycle, a sequencing polymerase binds an avidite to a polony and primer duplex, and traps a base-specific avidite to the polony. The result forms an extremely tight complex that enables a 100-fold reduction in reagent concentration compared to sequencing-by-synthesis (SBS).
AVITI and ABC reset expectations on quality scores (Q scores), delivering exceptional Q30 accuracy for 2 x 150 sequencing at > 90% and > 85% for 2 x 300 sequencing. AVITI demonstrated higher accuracy compared to legacy sequencing technology. Data shows fewer soft-clipped reads in difficult homopolymer and repeat regions.
*Note: Per Element recommendations we cannot run libraries that are more than 6 months old. Therefore, your library material will be disposed of 6 months after they are built in accordance with our SOPs.
Data Analysis: Supplementary data analysis will incur additional labor rates. Please contact Arizona Genetics Core to discuss further data analyis needs.
Price
Academic (UA): $varies| Academic (non-UA): $varies| Industry: $varies | Unit: Per Flow Cell Selected
Note- Flow cells are priced separately. Contact Arizona Genetics Core for a quote including library prep, sequencing and data analysis services.
Turnaround Time
Average turnaround time will vary depending upon application and sample input. Average data turnaround time for most NGS projects is approximately 4-8 weeks. The Arizona Genetics Core operates on a first come, first serve entry into sample queue.
Related Services
High Recovery DNA Extraction, RNA Extraction and analysis (Agilent Bioanalyzer 2100), Sanger DNA sequencing for validation of NGS data
Additional Information
If there is a specific application you are interested using the Element AVITI System but do not see listed here, please contact us to see if we can accommodate your needs.
University Customers: We are located in Keating Building Rm 106. Samples may be left in the freezer at the north entrance to our lab. Samples should be clearly labeled with your iLab Service ID.
External Customers: Samples should be shipped pre-frozen and preferably on dry ice (although regular ice packs work well enough for overnight shipping). Sample tubes should be sealed with parafilm.
Shipping Address:
Arizona Genetics Core
ATTN: NGS
1657 E. Helen Street
Keating Building, Rm 106
Tucson, AZ 85721
